Result of SNAPG: Difference between revisions
No edit summary |
|||
| Line 35: | Line 35: | ||
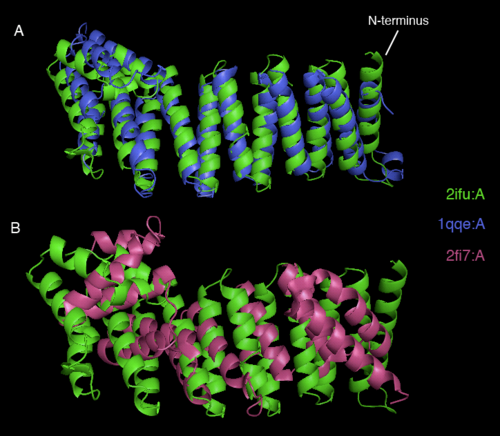
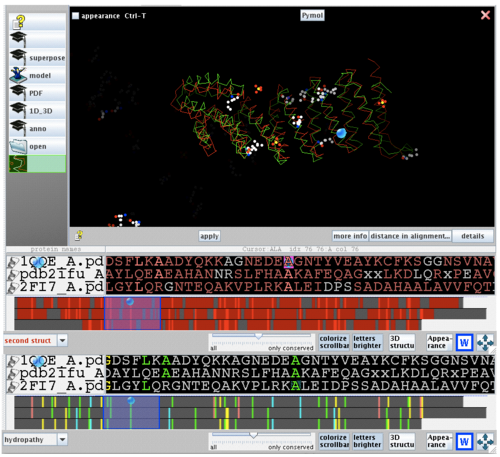
[[image:2ifu 1qqe 2fi7 alignment.png|thumb|left|500px|[[Media:2ifu 1qqe 2fi7 alignment.png|Figure 3. Stucture and sequence alignment of 2ifuA, 1qqeA and 2fi7A generated by STRAP. ]]]] | [[image:2ifu 1qqe 2fi7 alignment.png|thumb|left|500px|[[Media:2ifu 1qqe 2fi7 alignment.png|Figure 3. Stucture and sequence alignment of 2ifuA, 1qqeA and 2fi7A generated by STRAP. ]]]] | ||
| Line 40: | Line 41: | ||
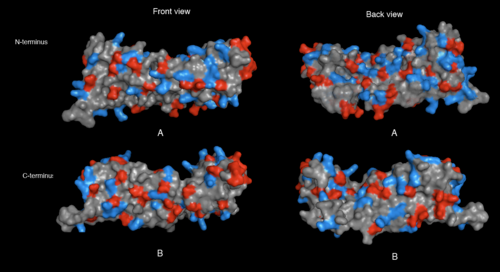
[[image:2ifuA-1qqe hydrophobes and hydrophilics.png|thumb|left|500px|[[Media:2ifuA-1qqe hydrophobes and hydrophilics.png|Figure. ]]]] | [[image:2ifuA-1qqe hydrophobes and hydrophilics.png|thumb|left|500px|[[Media:2ifuA-1qqe hydrophobes and hydrophilics.png|Figure 5. Hydrophobic and hydrophilic residues of 2ifuA (A) and 1qqeA (B) in front and back view. Amino acid residues with hydrophobic properties such as Ala, Gly, Val, Ile, Phe, and Met were selected and colored in green while that with hydrophilic properties (= Arg, Lys, His, Glu, Asp, Asn, Gln, Thr, Ser, and Cys) were colored in orange]]]] | ||
[[image:2ifuA-1qqe charged resi.png|thumb|left|500px|[[Media:2ifuA-1qqe charged resi.png|Figure. ]]]] | [[image:2ifuA-1qqe charged resi.png|thumb|left|500px|[[Media:2ifuA-1qqe charged resi.png|Figure. ]]]] | ||
Revision as of 06:01, 11 June 2007
SNAPG Structure
Structure architecture
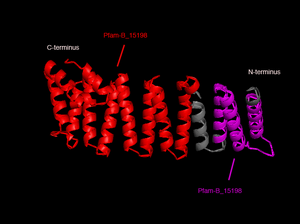
In order to analyze protein structure of SNAPG, structural comparison to known protein structure is required. An insight to SNAPG structural arrangement provides various informative data on possible protein functions and interactions with another protein and/or DNA. Based on protein families database, Pfam at Sanger, it was found that SNAPG protein matched to Pfam-B protein families and consist of 2 domains, Pfam-B_7270 (PB007270) and Pfam-B_15198 (PB015189) respectively as shown in Figure 1. Both of 2 domains appears to be associated with NSF attachment protein activity (*dijelasin di discussion aja! NSF acivity dijelasin ga?)[1].
. The sequence was also used against InterproScan generated by Profunc that gave an TFR (Tetratricopeptide-like helical) domain classification while that of protein family agrees with Pfam classification.
(interproscan)
Structural comparison
Dali webserver is one of the powerful tool to screen any protein that are structurally homologous with our query. Two structurally related proteins with highest Z-value generated by Dali server were chosen for SNAPG structure comparison analysis. These proteins were vesicular transport ptotein sec17 (1qqe) and type 4 fimbrial biogenesis protein (2f17) (refer to Table 1).alto
| PDB-chain | Structure | Z-value | % identity | Protein |
|---|---|---|---|---|
| 2IFU-A | 37.8 | 100 | Endocytosis/exocytosis. Gamma-SNAP (Danio rerio) | |
| 1QQE-A | 23.3 | 23 | Protein binding. Vesicular transport protein sec17(yeast) | |
| 2FI7-A | 12.9 | 14 | Protein transport. Type 4 fimbrial biogenesis protein pili (Pseudomonas aeruginosa) |
2ifuA, 1qqeA and 2fi7 alignment
physical properties