Methods 2ece: Difference between revisions
No edit summary |
No edit summary |
||
| Line 83: | Line 83: | ||
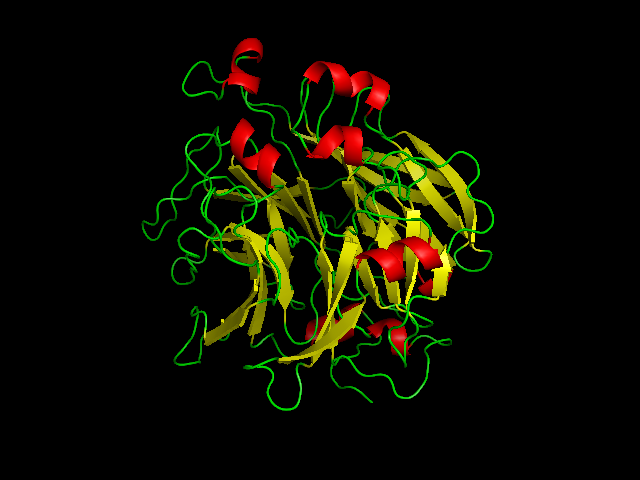
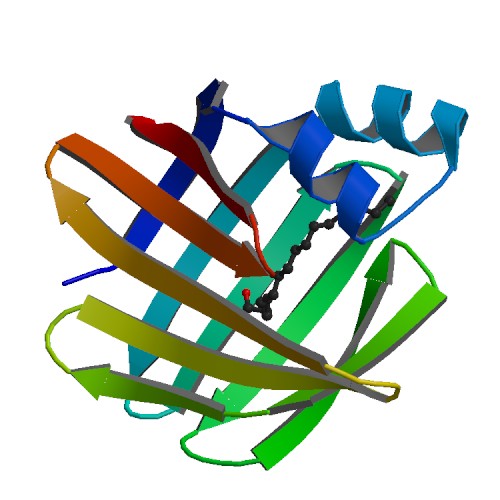
'''Figure 4''' Shows structural helices(red) and beta sheets (yellow) of SBP designed from pymol | '''Figure 4''' Shows structural helices(red) and beta sheets (yellow) of SBP designed from pymol | ||
[[Image:movie0001.png|frame|border|'''Figure 4''' Shows structural helices(red) and beta sheets (yellow) of SBP designed from pymol | |||
]] | |||
Revision as of 07:09, 9 June 2008
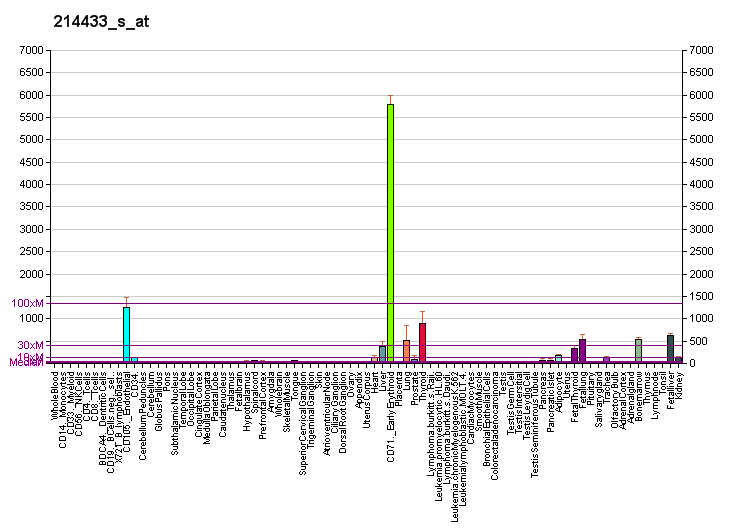
STRUCTURAL ANALYSIS
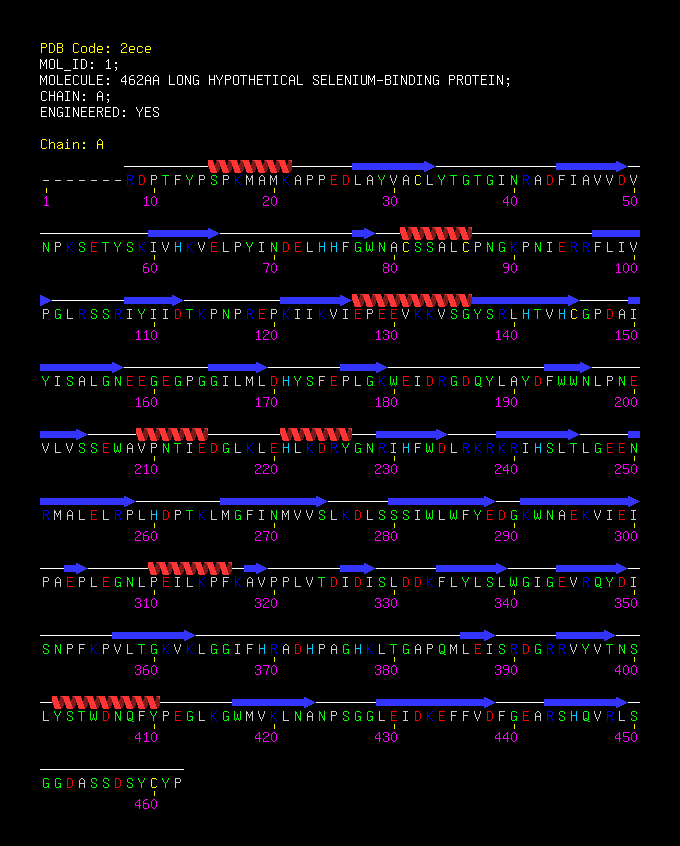
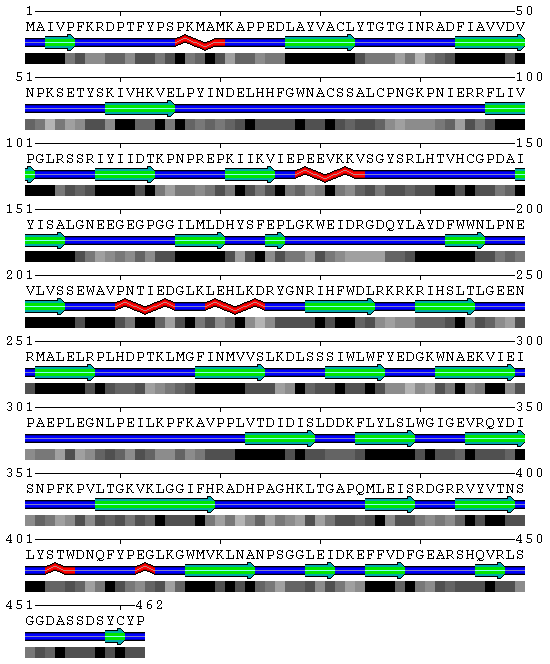
SBP 1 Amino Acid FASTA FORMAT Sequence
>2ECE:A|PDBID|CHAIN|SEQUENCE
MAIVPFKRDPTFYPSPKMAMKAPPEDLAYVACLYTGTGINRADFIAVVDVNPKSETYSKIVHKVELPYINDELHHFGWNA CSSALCPNGKPNIERRFLIVPGLRSSRIYIIDTKPNPREPKIIKVIEPEEVKKVSGYSRLHTVHCGPDAIYISALGNEEG EGPGGILMLDHYSFEPLGKWEIDRGDQYLAYDFWWNLPNEVLVSSEWAVPNTIEDGLKLEHLKDRYGNRIHFWDLRKRKR IHSLTLGEENRMALELRPLHDPTKLMGFINMVVSLKDLSSSIWLWFYEDGKWNAEKVIEIPAEPLEGNLPEILKPFKAVP PLVTDIDISLDDKFLYLSLWGIGEVRQYDISNPFKPVLTGKVKLGGIFHRADHPAGHKLTGAPQMLEISRDGRRVYVTNS LYSTWDNQFYPEGLKGWMVKLNANPSGGLEIDKEFFVDFGEARSHQVRLSGGDASSDSYCYP
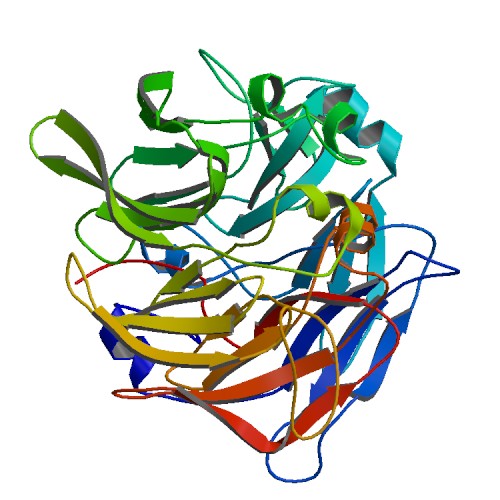
WARNING! Given sequence appeared to be a soluble protein, no TM domains found!
Output format is the following:
1st line -> residue numeration
2nd line -> query amino acid sequence
3rd line -> trans-membrane domain prediction (T-TM region, N-soluble part)
MAIVPFKRDPTFYPSPKMAMKAPPEDLAYVACLYTGTGINRADFIAVVDVNPKSETYSKI
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
VHKVELPYINDELHHFGWNACSSALCPNGKPNIERRFLIVPGLRSSRIYIIDTKPNPREP
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
KIIKVIEPEEVKKVSGYSRLHTVHCGPDAIYISALGNEEGEGPGGILMLDHYSFEPLGKW
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
EIDRGDQYLAYDFWWNLPNEVLVSSEWAVPNTIEDGLKLEHLKDRYGNRIHFWDLRKRKR
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
IHSLTLGEENRMALELRPLHDPTKLMGFINMVVSLKDLSSSIWLWFYEDGKWNAEKVIEI
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
PAEPLEGNLPEILKPFKAVPPLVTDIDISLDDKFLYLSLWGIGEVRQYDISNPFKPVLTG
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
KVKLGGIFHRADHPAGHKLTGAPQMLEISRDGRRVYVTNSLYSTWDNQFYPEGLKGWMVK
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
LNANPSGGLEIDKEFFVDFGEARSHQVRLSGGDASSDSYCYP
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
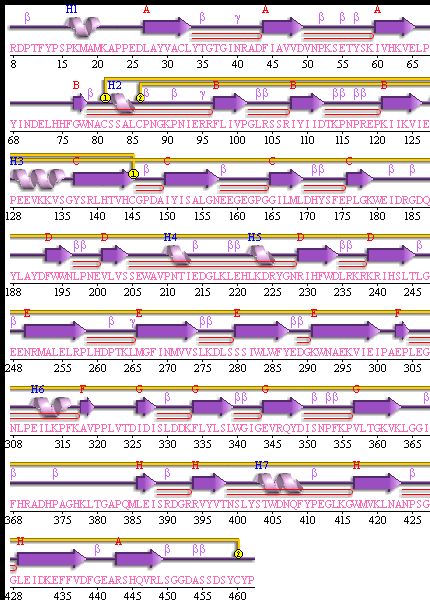
Figure 3 SABLE server results.
Figure 4 Shows structural helices(red) and beta sheets (yellow) of SBP designed from pymol
Figure 5
Figure 6
STRUCTURAL COMPARISONS
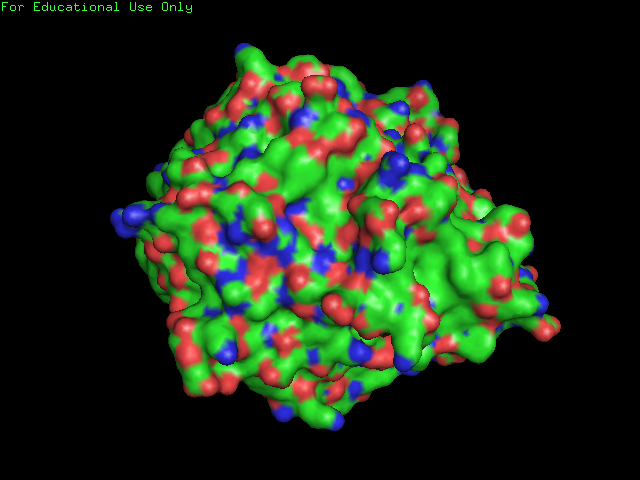
Surface Structure of SBP: Note the few clefts compared to the DNA isomerase shown below.
Rat fatty acid binding protein 2IFB, a was found to have 92.5% homology to 14 KDa Selenium binding protein purified from rat liver using column chromatography and SDS-Gel techniques.([ref 3])
Explore SBP features and structural summary here [2].The domains of SBP are shown here [3] Notice how the domains are similar to the putative Isomerase domains of E.coli below.
1RI6 DOMAINS
2ECE DOMAINS
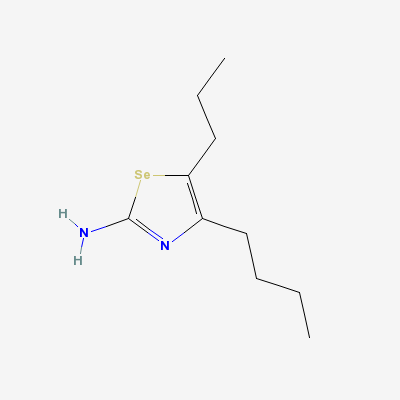
(Complex Of Bovine Odorant Binding Protein (Obp) With A Selenium Containing Odorant)"Image:Ligand of bovine.png" [[4]]
SEQUENCE ANALYSIS
Selenium binding protein 1 (SELENBP1) SELECTED PROTEIN SIMILARITIES Comparison of sequences in UniGene with selected protein reference sequences. The alignments can suggest function of a gene. [5]