Arylformamidase Results
Structure
2PBL Biological Structure
The functional biological structure of 2PBL is assumed by PDB to be a monomer (see figure 3) even though the 'whole' protein is shown to be interacting with chains A, B, C and D.
2PBL Structural Similarity
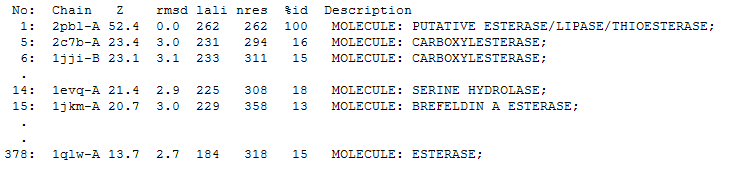
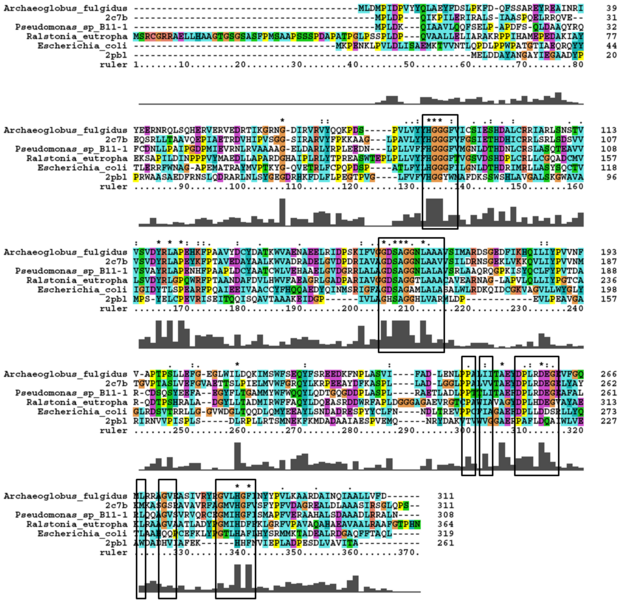
The DALI tool produces proteins that are structurally similar to the protein of interest. The search result showed similarities to be mostly carboxylesterases/hydrolases. The first significant hit from DALI was a carboxylesterase from a metagenomic Archeaon. The second significant hit was a carboxylesterase of Archaeoglobus fulgidus. Analysis of 2PBL secondary structure similarity with the Archeal carboxylesterases showed conservation of the order of occurrence of different conformational types. For instance in all three proteins, the first conformation type is a helix and then three beta strands followed by a helix (see figures 4 & 5).
The catalytic triad structure
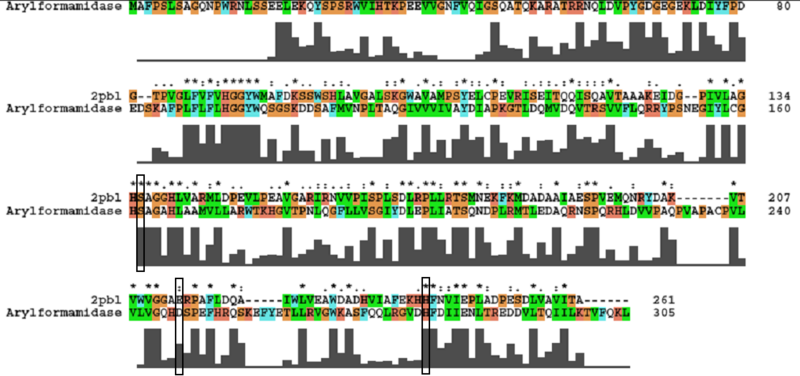
A number of carboxylesterases perform their hydrolysis function using specific catalytic residues. The ClustalW alignment showed conservation of the Archeal carboxylesterases catalytic triad in 2PBL. The residues are Serine (Ser) 137, Glutamate (Glu) 215 and Histidine (His) 242. Residues Ser and His were fully conserved whereas E was semi-conserved. Using the human arylformamidase it was observed that Aspartate (asp) was used for eukaryotes instead of Glu, which is used for prokaryotes.
Residues that were highly conserved between prokaryotic and eukaryotic species were annotated on the structure of 2PBL (see figure...). Distances between the conserved residues of the catalytic triad were measured. The blue region in the figure below shows the clustering of conserved residues around the unknown ligand.
Conserved - Asp53, His69, Gly70, Gly71, Trp73, Gly134, Ser136, Ala137, Gly138, His241. Semi-conserved - Tyr72, His140, Ser166, Leu168, Leu171, Leu174, Glu214, Val244, Leu248. Catalytic triad - Ser136, Glu214, His241.
Function
The most similar sequence from the BLAST search with functional information available was an arylformamidase isolated from the liver of Mus musculus (see figure 8). A functional analysis of this arylformamidase has been performed identifying a catalytic triad using site-directed mutagenesis (Pabarcus et al. 2007). Conservation of this catalytic triad with 2PBL was assessed using sequence homology (see figure 9). Both residues Ser162 and His279 were found to be identical in relatively conserved regions of the alignment. However, Asp247 had undergone a semi-conservative substitution to glutamic acid. These residues correlated to Ser136, Glu214 and His241 of 2PBL which were subsequently located on the tertiary structure and determined to be sufficiently proximal to one another for catalysis (see figure 10).
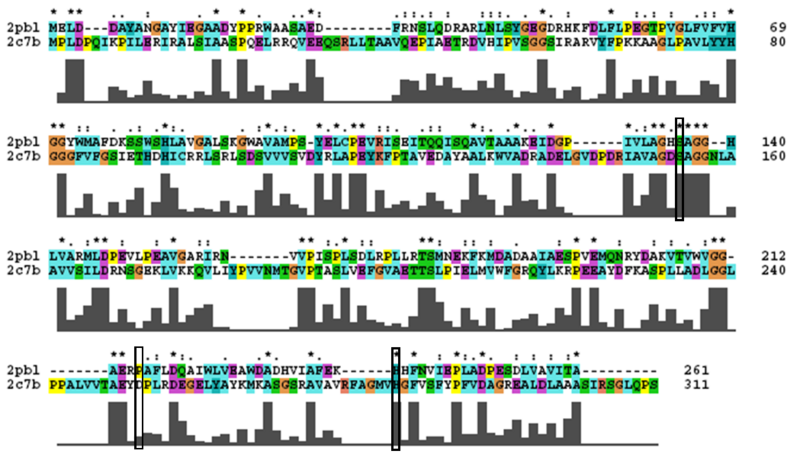
2PBL was found to share most structural similarity with a thermostable carboxylesterase from a metagenomic archaeon (PDB ID: 2C7B; see figure 11). 2C7B shares a 16% sequence identity with 2PBL. From its structure, a catalytic triad has been identified with the residues Ser154, Asp251 and His281 (Byun, et al. 2007). To substantiate any functional similarity between 2PBL and 2C7B, structural conservation of the 2C7B catalytic triad was analysed (see figure 12). Both Ser154 and His281 matched, but the aspartic acid had undergone a semi-conservative substitution to glutamic acid.
A number of proteins from the hormone-sensitive lipase (HSL) class of lipolytic enzymes as identified by Byun, et al. (2007) was found within top-scoring results of the DALI search (see figure 11). To characterise sequence similarity, a multiple sequence alignment of the amino acid sequences for these structures was performed (see figure 12).
Sequence & Homology
The phylogenetic tree constructed using the multiple sequence alignment on 2PBL related sequences is shown in figure .... The bootstrap values reveal low confidence with many of the nodes occurring lower down on the phylogenetic tree. This reveals a possible explanation for certain closely related species to be grouped into separate clades. Despite low bootstrap scores, the grouping reliably separates prokaryotes from eukaryotes. Interestingly, homologous eukaryotes include yeasts and moulds, plants, invertebrates and vertebrates.
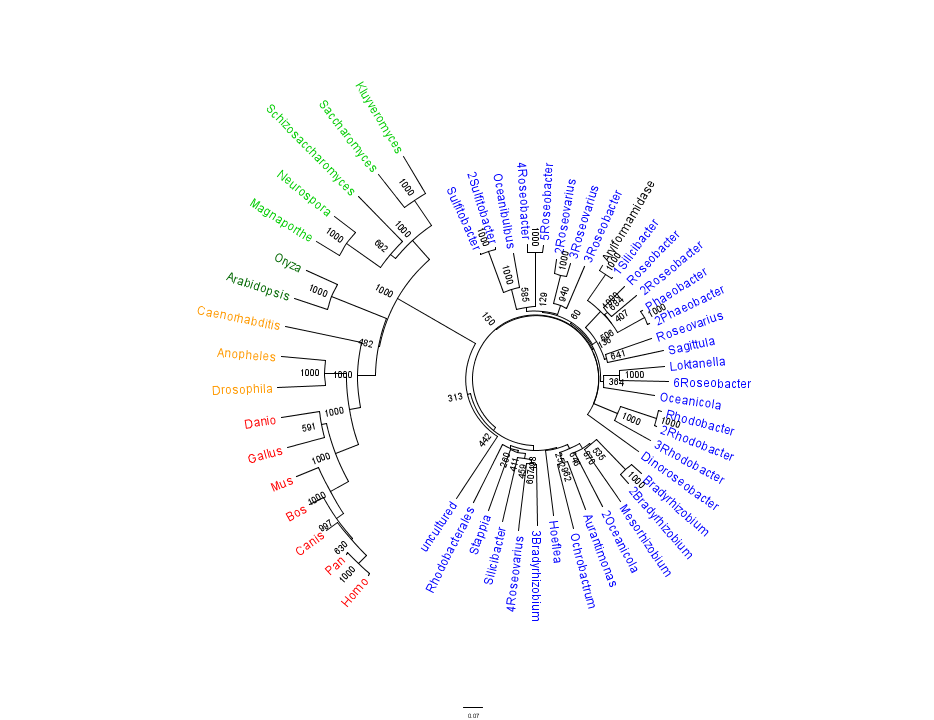
To further elucidate the phylogeny of 2pbl, its human homologue, an arylformamidase, was queried in a BLAST search. The top scoring matches of bacterial homologues, present in figure 1, were appended with top scoring matches of eukaryotic homologues. The human homologue has a 26.28% sequence similarity. Despite this low score, multiple sequence alignment revealed that key regions were highly conserved between bacterial and eukaryotic homologues. This was demonstrated in a phylogenetic analysis of 2PBL and its human homologue which was largely consistent with traditional taxonomic groupings of organisms(see figure ...). Specifically, it reveals greater statistical confidence in the separation of prokaryotes and eukaryotes.
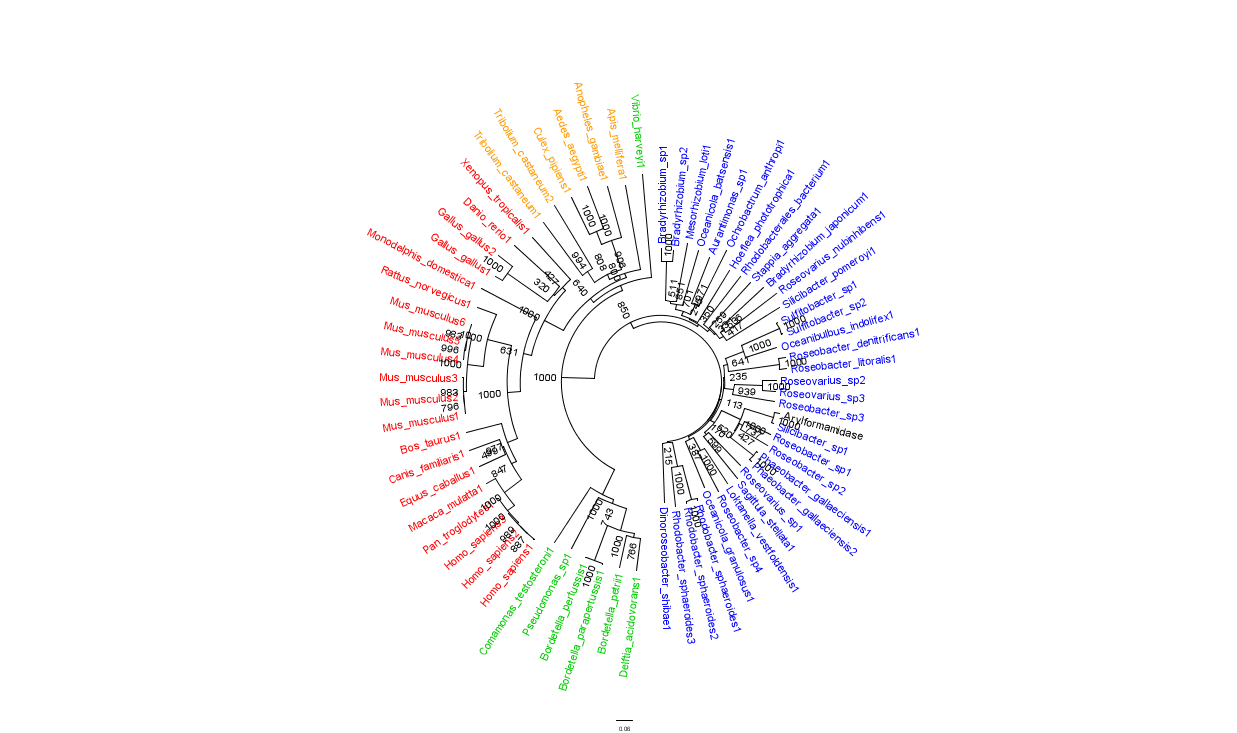
In general, members of the same genus have been grouped together on these phylogenetic trees with some notable exceptions. Notably, members of the Silicibacter clade occur on disparate branches of the tree.